Note: See also these related newsletters: 20 Feb 2020 (link) on DTR as part of the language of resistance and 7 June 2020 (link) on using DTR to estimate antibiotic value. There is also a 5-minute DTR explainer on YouTube that you might find useful.
Dear All:
When thinking about studies of resistant pathogens, issues of case-finding and statistical design are frequent topics of debate. As an example, the recent De Kraker et al. (CID 2018) paper by the COMBACTESTAT-Net group gives a great survey of ideas for optimizing design and analysis of these trials. (Note that full titles and cites of all papers mentioned in this note are given below my signature.)
A less-discussed area of late (at least to my ear) has been the implications of the categories of resistance. Based on expert consensus, CDC and ECDC gave us a widely used set of ideas in Magiorakos et al. (CMI 2012):
- MDR (Multi-Drug Resistance) was defined as acquired non-susceptibility to at least one agent in three or more antimicrobial categories.
- XDR (eXtensive Drug Resistance) was defined as non-susceptibility to at least one agent in all but two or fewer antimicrobial categories (i.e., bacterial isolates remain susceptible to only one or two categories).
- PDR (Pan Drug Resistance) was defined as non-susceptibility to all agents in all antimicrobial categories.
McDonnell et al. (disclaimer: I am an author on both papers mentioned in this paragraph) added to this list by proposing (CID 2016) the idea of UDR (Usual Drug Resistance) to describe isolates that aren’t fully susceptible wild-type strains but that can nonetheless be readily treated with standard therapies. UDR is useful shorthand for the setting in which a novel agent is tested against an existing standard-of-care agent in a non-inferiority trial (for more on this, see the expanded discussion in CID 2017).
Now adding to this conversation, Kadri et al. (CID 2018) propose the idea of DTR: Difficult-to-Treat Resistance. Their point of departure is the idea that the MDR-XDR-PDR definitions make no distinction between strengths and weaknesses of the individual antibiotics: agents with higher efficacy and lower toxicity are counted in the same way as agents with lower efficacy and higher toxicity.
To address this, Kadri et al. define DTR as resistance to all of the typical first-line, lower toxicity agents, defined by Kadri et al. as the beta-lactams (including carbapenems and beta-lactamase inhibitor combinations) and the fluoroquinolones. In this setting, agents with toxicity or pharmacology limitations (e.g., the nephrotoxic aminoglycosides) must be used.
9 Apr 2026 addendum: Visually, Kadri 2018 includes this graphic showing the idea of DTR … note how DTR include all of PDR but only portions of MDR and XDR:
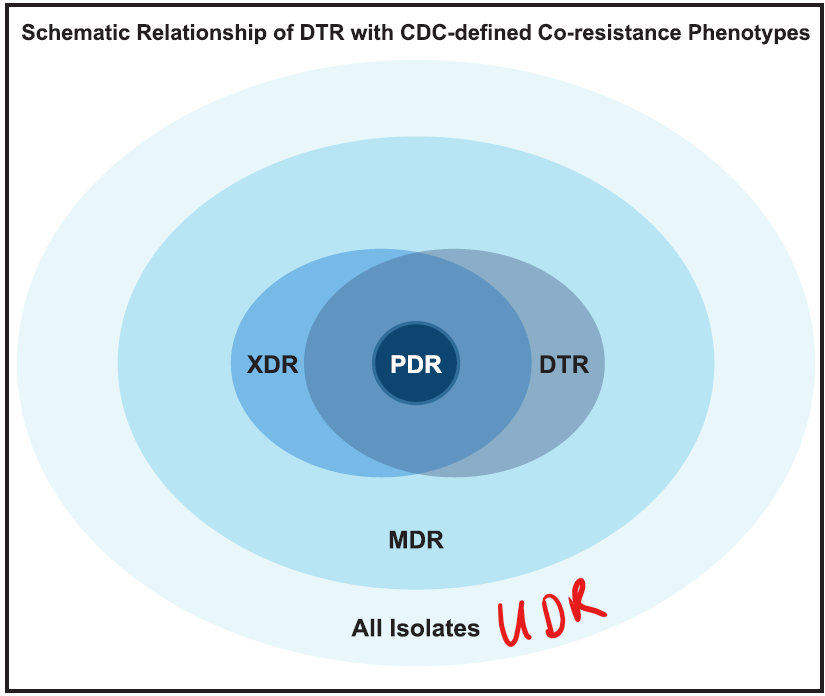
Kadri et al. argue that discriminating among the qualities of the agents that must be used as resistance increases may help explain why Lye et al. (CMI 2012) found that “multidrug resistance per se is not associated with higher mortality when effective antibiotics are used for definitive therapy.” As another example, this might help explain in part the high and similar mortality with both colistin monotherapy and colistin-based combination therapies in the prospective study reported by Paul et al. (LancetID 2018, see also the related editorial by Perez and Bonomo, and a helpful secondary analysis by Zak-Doron and colleagues).
And, their thinking proved correct. Based on data from the Premier Healthcare Database from 173 US institutions on 46,521 Gram-Negative Blood Stream Infection (GNBSI) isolates from the period 2009-2013, about 1.5% of isolates were DTR. Critically, patients with DTR GNBSI isolates had a 40% higher adjusted mortality risk than those with non-DTR GNBSI. These findings were consistent in a variety of sensitivity analyses including by-species analyses.
This is a fascinating and helpful result and it is worth the effort to work carefully through the entire Kadri paper as well as the related papers mentioned above. As a simple but very meaningful concept that can be readily applied at the bedside, I think we’re going to see DTR used repeatedly as we go forward.
All best wishes, –jr
John H. Rex, MD | Chief Medical Officer, F2G Ltd. | Expert-in-Residence, Wellcome Trust. Follow me on Twitter: @JohnRex_NewAbx. See past newsletters and subscribe for the future: https://amr.solutions/blog/
Full titles and cites of papers
- de Kraker, M. E. A., H. Sommer, F. de Velde, I. Gravestock, E. Weiss, A. McAleenan, S. Nikolakopoulos, O. Amit, T. Ashton, J. Beyersmann, L. Held, A. M. Lovering, A. P. MacGowan, J. W. Mouton, J. F. Timsit, D. Wilson, M. Wolkewitz, E. Bettiol, A. Dane, S. Harbarth and C.-N. Consortium (2018). “Optimizing the Design and Analysis of Clinical Trials for Antibacterials Against Multidrug-resistant Organisms: A White Paper From COMBACTE’s STAT-Net.” Clin Infect Dis 67(12): 1922-1931. (link)
- Kadri, S. S., J. Adjemian, Y. L. Lai, A. B. Spaulding, E. Ricotta, D. R. Prevots, T. N. Palmore, C. Rhee, M. Klompas, J. P. Dekker, J. H. Powers, 3rd, A. F. Suffredini, D. C. Hooper, S. Fridkin, R. L. Danner and I. National Institutes of Health Antimicrobial Resistance Outcomes Research (2018). “Difficult-to-Treat Resistance in Gram-negative Bacteremia at 173 US Hospitals: Retrospective Cohort Analysis of Prevalence, Predictors, and Outcome of Resistance to All First-line Agents.” Clin Infect Dis 67(12): 1803-1814. (link)
- Lye, D. C., A. Earnest, M. L. Ling, T. E. Lee, H. C. Yong, D. A. Fisher, P. Krishnan and L. Y. Hsu (2012). “The impact of multidrug resistance in healthcare-associated and nosocomial Gram-negative bacteraemia on mortality and length of stay: cohort study.” Clin Microbiol Infect 18(5): 502-508. (link)
- Magiorakos, A. P., A. Srinivasan, R. B. Carey, Y. Carmeli, M. E. Falagas, C. G. Giske, S. Harbarth, J. F. Hindler, G. Kahlmeter, B. Olsson-Liljequist, D. L. Paterson, L. B. Rice, J. Stelling, M. J. Struelens, A. Vatopoulos, J. T. Weber and D. L. Monnet (2012). “Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance.” Clin Microbiol Infect 18(3): 268-281. (link)
- McDonnell, A. M., J. H. Rex, H. Goossens, M. Bonten, V. G. Fowler and A. Dane (2016). “Efficient Delivery of Investigational Antibacterial Agents via Sustainable Clinical Trial Networks.” Clinical Infectious Diseases 63 (Suppl. 2): S57-S59. (link)
- Paul, M., G. L. Daikos, E. Durante-Mangoni, D. Yahav, Y. Carmeli, Y. D. Benattar, A. Skiada, R. Andini, N. Eliakim-Raz, A. Nutman, O. Zusman, A. Antoniadou, P. C. Pafundi, A. Adler, Y. Dickstein, I. Pavleas, R. Zampino, V. Daitch, R. Bitterman, H. Zayyad, F. Koppel, I. Levi, T. Babich, L. E. Friberg, J. W. Mouton, U. Theuretzbacher and L. Leibovici (2018). “Colistin alone versus colistin plus meropenem for treatment of severe infections caused by carbapenem-resistant Gram-negative bacteria: an open-label, randomised controlled trial.” Lancet Infect Dis 18(4): 391-400. (link)
- Perez, F. and R. A. Bonomo (2018). “Evidence to improve the treatment of infections caused by carbapenem-resistant Gram-negative bacteria.” Lancet Infect Dis 18(4): 358-360. (link)
- Rex, J. H., G. H. Talbot, M. J. Goldberger, B. I. Eisenstein, R. M. Echols, J. F. Tomayko, M. N. Dudley and A. Dane (2017). “Progress in the fight against multidrug-resistant bacteria 2005-2016: Modern non-inferiority trial designs enable antibiotic development in advance of epidemic bacterial resistance.” Clinical Infectious Diseases 65: 141-146. (link)
- Zak-Doron, Y., Y. Dishon Benattar, I. Pfeffer, G. L. Daikos, A. Skiada, A. Antoniadou, E. Durante-Mangoni, R. Andini, G. Cavezza, L. Leibovici, D. Yahav, N. Eliakim-Raz, Y. Carmeli, A. Nutman, M. Paul and A. S. Group (2018). “The Association Between Empirical Antibiotic Treatment and Mortality in Severe Infections Caused by Carbapenem-resistant Gram-negative Bacteria: A Prospective Study.” Clin Infect Dis 67(12): 1815-1823. (link)
Upcoming meetings of interest to the AMR community:
- 15 Jan 2018 (London): BSAC’s Antimicrobial Chemotherapy Conference 2019: “An ABC for everyone involved in developing new antimicrobials.” Details here.
- 29 Jan 2018 (REVIVE webinar): “Clinical development for non-developers Part 3: Antibacterial Drug Enhancer Combinations and Non-traditional Products.” Register here.
- 4-5 Feb 2019 (London): Hamied Foundation UK-India Antimicrobial Resistance Meeting 2019. This is a 2-day meeting focused on building research links between the UK and India with the specific aim of jointly addressing the challenge of AMR. Register here.
- 14-15 Mar 2019 (Berlin): BEAM-, CARB-X-, Novo REPAIR-, DZIF-, ND4BB-ENABLE-sponsored (among a long list!) Berlin Conference on Novel Antimicrobials and AMR Diagnostics. Details here. Poster submissions are being accepted through 9 Jan (details here).
- 21-22 Mar 2019 (Birmingham, UK): BSAC Spring Conference.
- 26 Mar 2019 (London, UK): Sponsored by The Economist, a 1-day symposium entitled “Antimicrobial Resistance: Preventing an antibiotic apocalypse.” Register here.
- 13-16 Apr 2019 (Amsterdam): Annual ECCMID meeting
- 16-18 Apr 2019 (Utrecht): ICOHAR, International Conference on One Health Antimicrobial Resistance. Organized by the ESCMID Study Group for Veterinary Microbiology (ESGVM).
- 24-26 Apr 2019 (Boston): Annual SHEA (Soc. for Hospital Epidemiology of America) Spring meeting
- 6-11 May 2019 (Ljubljana, Slovenia): 37th Annual Meeting of the European Society for Paediatric Infectious Diseases (ESPID). Details here.
- 3-6 Jun 2019 (Philadelphia): Annual BIO meeting
- [NEW] 10-11 June 2019 (Research Triangle Park, NC): AMR Action Summit on R&D and Commercialization. Sponsors include the British-American Business Council, the UK Gov’t, CARB-X, the NC Biotechnology Center, and others. Details here.
- 20-24 June 2019 (San Francisco): Annual ASM Microbe meeting.
- [Mark your calendar now!] 3-6 Sep 2019 (Boston). Annual ASM-ESCMID Conference on Antibiotic Development. The Bootcamp series will continue on 3 Sep with the main meeting on 4-6 Sep. Mark your calendar now and check back here for details.
- 6-8 Sep 2019 (Bilbao, Spain): 5th ESCMID conference on Vaccines. Check back here for details.
- 2-6 Oct 2018 (Washington, DC): IDSA’s annual IDWeek meeting.
- 19-27 Oct 2019 (Annecy, France): International Course on Antibiotics and Resistance (ICARe) – A soup-to-nuts intensive residential training program on all things AMR, especially R&D for new antibiotics. See this link for details.