Dear All,
Building on the ATLAS database (link) that provides open access to data from 2004 to the present out of the multiple surveillance programs that Pfizer has run over that interval, Wellcome Trust and Pfizer have just announced that the underlying database will also be used to support SPIDAAR (Surveillance Partnership to Improve Data for Action on Antimicrobial Resistance), a new multi-year, public-private research collaboration with the governments of Ghana, Kenya, Malawi and Uganda to track resistance patterns and better understand the burden of antimicrobial resistance (AMR) on patients living in low- and middle-income countries.
To decode this, let’s start with some background on ATLAS. Pfizer has had a number of marketed antibacterial and antifungal agents over the past 20 years (tigecycline, ceftaroline, linezolid, fluconazole, and voriconazole … just to name a few). Triggered by conversations on the importance of sharing data on AMR (see, for example, this 2018 workshop at Wellcome trust, link), Pfizer created an open-access database in 2017 that provides access to data from all the surveillance programs that Pfizer has variously run since 2004 (link to an early press release on ATLAS; link to general current description of the project; link to ATLAS itself) .
The scope of the data in the database is impressive: there are MICs on 680k bacterial isolates and 45 antibacterials from 76 countries as well as MICs on 15.5k fungi (mostly yeasts) and 10 antifungals from 40 countries. The database is very interactive and a great resource for graphics. Here, for example, is the distribution of carbapenem-resistant P. aeruginosa in 2004 vs. 2018:
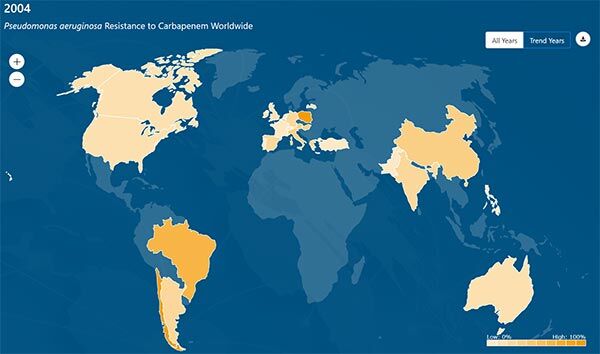
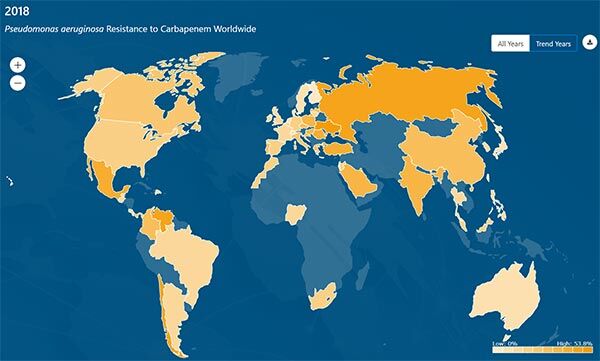
You can also get the database to play back the timeline of the spread of resistance: after picking the bugs, drugs, and years, you just press play to watch the resistance creep across the globe! A little scary but also very instructive! I encourage you to have a look (and see below for more … you can download the entire 121MB database from the AMR Register).
—
So, what about SPIDAAR? As noted above, this is collaboration between Wellcome Trust, Pfizer, and the governments of Ghana, Kenya, Malawi, and Uganda to build capacity by initiating a surveillance program in these countries. Data from this work would obviously have value for each country’s National Action Plan on AMR.
As noted above, SPIDAAR will use the ATLAS database as a vehicle for sharing its data. The SPIDAAR program also includes a separate real-world data study to examine the impact of antibiotic resistance on clinical as well as economic outcomes by recruiting a prospective cohort of patients with hospital-acquired infections.
As noted in an online article about the project by Wellcome Trust’s Gemma Buckland-Merrett (link), surveillance data is important at both the global (e.g., setting priorities) and local (e.g., guiding targeted action) level in efforts vs AMR. Indeed, The Access to Medicine Foundation recently called out the ATLAS project itself by noting (link) that through ATLAS, Pfizer was “… the first company to share raw data on the spread of resistance so that third parties can explore the potential for further research.”
This is really excellent to see! To my eye, this is a first of its kind public-private partnership supporting surveillance in countries with limited capacity and capability and is proof of how multiple sectors can come together to address AMR.
More broadly, this is the type of initiative called for by the AMR Industry Alliance (AMR IA) in its reports on Industry commitments (link) to “Support collaboration and sharing of relevant non-proprietary data” and “Collect and share surveillance data.”
—
And, what does this mean for you? Well, ATLAS is one of the programs feeding in to the AMR Research Initiative (aka, the AMR Register), a Wellcome Trust-funded initiative (link). The AMR register website currently shows 13 surveillance programs where data are available either by request (12 of the 13) or by direct download (ATLAS: go here for descriptive webpage and here for the 121 MB (!!) Excel download).
The goal of all of this is to encourage creative re-use of these data. An example of this was noted in the 25 April 2019 announcement by Wellcome of the winners of their recent Data Re-Use Prize (link to general webpage on the prize). One of the two prizes went to a team who used the ATLAS dataset to derive a composite index of antibiotic resistance for common infection syndromes with the aim of creating a guide to empirical therapy (link). Although the authors concluded that there were not enough isolates to fully inform their model, the strategy is certainly sound and shows how extended data collection (e.g., SPIDAAR) could enable such goals.
Wow! Mighty oaks from little acorns grow … and I look forward to watching these programs progress as others add data.
All best wishes, –jr
John H. Rex, MD | Chief Medical Officer, F2G Ltd. | Operating Partner, Advent Life Sciences. Follow me on Twitter: @JohnRex_NewAbx. See past newsletters and subscribe for the future: https://amr.solutions/blog/. All opinions are my own.
Current funding opportunities:
- Novo REPAIR Impact Fund is open for global applications through 31 Jul 2020. Go here for current details.
- 2020 funding rounds for CARB-X have not been announced.
- The Global AMR R&D Hub’s dynamic dashboard (link) summarizes funders and projects by geography, stage, and more.
Upcoming meetings of interest to the AMR community:
- 30 Jun 2020 (online, 17:00-18:30 CEST): GARDP REVIVE webinar. Title: “Clinical development of antimicrobials – Phase 1 development challenges.” Speaker: Markus Zeitlinger. Go here to register.
- 9 Jul 2020 (online, 09:00-10:30 CEST): GARDP REVIVE webinar. Title: “The challenges and opportunities for antimicrobial R&D in low- and middle-income countries – India case study.” Speaker: Anand Anandkumar and Kamini Walla. Go here to register.
- 17 Jul-2 Aug 2020 (Marine Biology Laboratory, Woods Hole, MA): Residential course entitled “Molecular Mycology: Current Approaches to Fungal Pathogenesis.” This 2-week intensive training program has run annually for many years and gets outstanding reviews. Go here for details.
- 27 Jul-31 Jul 2020 (online): Small World Initiative Instructor Training Workshop – training for undergraduate professors and high school teachers in wet lab techniques, parallel curricula, & pedagogical instruction to engage students in the hunt to find new antibiotics in soil (also covering distancing learning options). Go here to register.
- 4 Aug 2020 (Silver Spring): FDA workshop entitled “Development Considerations of Antifungal Drugs to Address Unmet Medical Need.” Go here to register.
- 5 Aug 2020 (Silver Spring): FDA workshop entitled “Developing Antifungal Drugs for the Treatment of Coccidioidomycosis (Valley Fever) Infection.” Go here to register.
- September 2020. University of Sheffield (UK). Applications are being taken for a new 1-year (full-time) or 2-year (part-time) Masters of Science course in Antimicrobial Resistance. The program runs annually from September and covers microbiology, clinical practice and policy. The course webpage is here.
- 9-10 Sep 2020 (Washington, DC): US PACCARB public meeting. Go here for details.
- 26-29 Oct 2020 (online meeting), Annual ESPID meeting (European Society for Pediatric ID, #38)
- 10-13 Apr 2021 (Vienna): Annual ECCMID meeting (#31)
- 20-24 June 2021 (Toronto): International Symposium on Pneumococci and Pneumococcal Diseases (ISPPD-12). Go here for details.
- 3-7 Jun 2021 (Anaheim), ASM Microbe 2021. Go here for details.
- 8-11 Oct 2021 (Aberdeen, Scotland): 10th Trends in Medical Mycology. Go here for details.
- 16-24 Oct 2021 (Annecy, France): Interdisciplinary Course on Antibiotics and Resistance (ICARe). This is a soup-to-nuts residential course on antibiotics, antibiotic resistance, and antibiotic R&D. The course is very intense, very detailed, and gets rave reviews. Registration is here and is limited to 40 students.
- 18-21 May 2021 (Albuquerque, New Mexico): Biannual meeting of the MSGERC (Mycoses Study Group Education and Research Consortium). Save-the-date announcement is here, details to follow.