Dear All (and with thanks to Patricia Bradford for co-authoring this newsletter),
When you are seeking novelty, antibacterial discovery is hard … really, really, REALLY hard. And it gets even harder if you want activity vs. Gram-negative bacteria. As the latest proof of this, a paper from GARDP’s Blasco et al. describing use of an HTS (High-Throughput Screen) shows us (again) how hard this is. Here are the links you need:
- [the new paper] Blasco, Jang, Terauchi, Kobayashi, Suzuki, Akao, et al. “High-throughput screening of small-molecules libraries identified antibacterials against clinically relevant multidrug-resistant A. baumannii and K. pneumoniae.” EBioMedicine. 2024 Mar 22;102:105073. https://doi.org/10.1016/j.ebiom.2024.105073.
- [overview of the general problem and of the results of 65 HTS campaigns at AZ] Perros M. “Infectious disease. A sustainable model for antibiotics.” Science. 2015 Mar 6;347(6226):1062-4. https://doi.org/10.1126/science.aaa3048.
- [deep dive into the data on 65 HTS campaigns at AZ] Tommasi, Brown, Walkup, Manchester, and Miller. “ESKAPEing the labyrinth of antibacterial discovery.” Nat Rev Drug Discov. 2015 Aug;14(8):529-42. https://doi.org/10.1038/nrd4572.
- [important prior paper reporting on 70 HTS campaigns at GSK] Payne, Gwynn, Holmes, and Pompliano. “Drugs for bad bugs: confronting the challenges of antibacterial discovery.” Nat Rev Drug Discov. 2007 Jan;6(1):29-40. https://doi.org/10.1038/nrd2201.
- [great general survey of discovery challenges] McDowell, Quinn, Leeds, Silverman, and Silver. “Perspective on Antibacterial Lead Identification Challenges and the Role of Hypothesis-Driven Strategies.” SLAS DISCOVERY: Advancing the Science of Drug Discovery. 2019;24(4):440-456. https://doi.org/10.1177/2472555218818786.
The paper from Blasco et al. is readily summarized:
- 48,015 small molecule compounds contributed by colleagues at Takeda, Eisai, and Daiichi Sakyo (thank you!!) were selected for screening. Of these, the compounds from Takeda and Eisai (38,415 compounds) were selected based on current understanding of physicochemical parameters (the “eNTRy rules”) suggesting the compounds would enter and then be retained in Gram-negative bacteria.
- Important point: Those 48,015 compounds were not just a random collection of molecules! Rather, they were a selected and curated subset of larger compound libraries at the companies — the authors sought to increase their chance of success before doing the HTS by applying best current knowledge (for the rules used, see Richter et al. from 2017 & 2018: https://doi.org/10.1038/nature22308 and https://doi.org/10.1111/nyas.13598).
- Using a whole cell screen (live bacteria in suspension tested in an HTS using the principles of EUCAST susceptibility methods to assay individual compounds in 384-well plates), the compounds were screened for activity vs. XDR Acinetobacter baumannii and MDR Klebsiella pneumoniae. See the 13 Jan 2019 newsletter entitled “Categories Of Resistance: MDR, XDR, PDR, UDR, And (New!) DTR” if you need a refresher on the ideas of XDR and MDR.
- Drum roll…
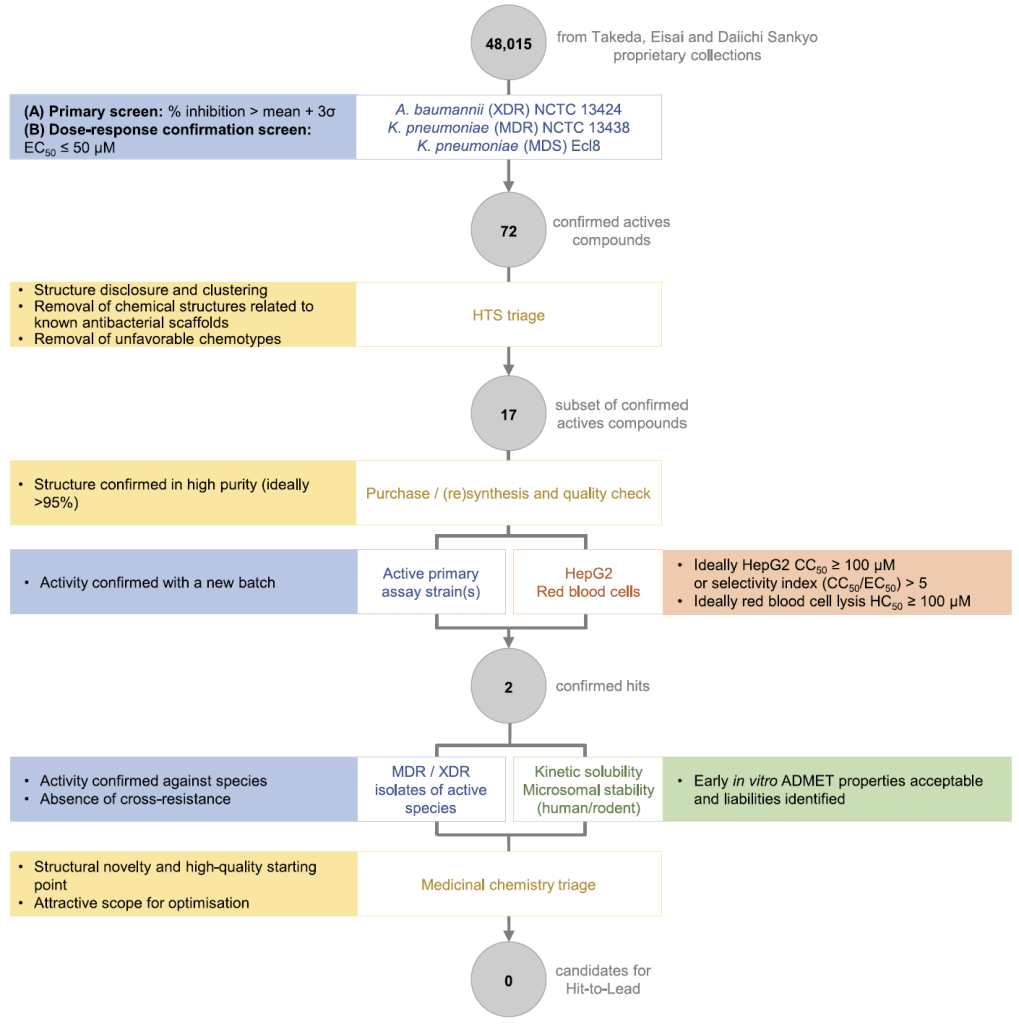
Yup … 48,015 curated compounds at the start, two confirmed hits, but in the end none with properties suggesting that the compounds were leads (that is, none that looked as if it could be improved by further work).
Unfortunately, it is not only HTS campaigns that have been disappointing with regards to an actual drug being discovered:
- State-of-the-art crystal structure docking was used by companies for rational drug design, again with no success with regards to identifying a clinical candidate (Franceschi and Duffy. “Structure-based drug design meets the ribosome.” Biochem Pharmacol. 2006 Mar 30;71:1016-25. https://doi.org/10.1016/j.bcp.2005.12.026).
- An AI-based approach reported in 2020 by Stokes et al. in Nature (https://doi.org/10.1016/j.cell.2020.01.021) led to excitement about a compound (halicin) that had broad-spectrium in vitro activity but that did not have the other properties needed to make a compound a drug. See this 5-part newsletter series (21 Feb 2020 et seq.) for an extended discussion of halicin, AI-based approaches, and further work from the group behind the paper on halicin.
- A natural product screen described in a 2015 paper in Nature by Ling et al. (https://doi.org/10.1038/nature14098) reported the discovery of texiobactin by using an innovative approach (the “iChip”) to growing otherwise uncultivatable bacteria. Texiobactin has a novel mechanism and in vitro activity vs. Gram-positives as well as Mycobacterium tuberculosis, but this product has also not advanced in the clinic.
Finally, the paucity of progressible compounds coming out of any kind of screening effort is further highlighted in two recent reviews of the area.
- In a 2023 summary of antibiotics approved since 2000 (Shi et al., https://doi.org/10.3390/molecules28041762), only bedaquiline was discovered using a screen.
- In a 2024 review of anti-Gram-negative agents in clinical development (Paterson, https://pubmed.ncbi.nlm.nih.gov/38445383/), none of the compounds listed as being in Phase 2 or Phase 3 represent a novel chemotype. A few of the compounds listed in Phase 1 were identified via HTS, however it is too soon to know if these will progress into efficacy studies.
Deep sigh. We knew that discovery was hard, but these papers really drive home the painful truth that it is reasonably easy to find compounds that kill bacteria but finding well-behaved (drug-like) molecules that kill bacteria is very, very, very, very hard. As summarized by Kevin Outterson (personal communication), we need approximately 6,000 basic research projects per decade to yield 6 high-impact treatments:
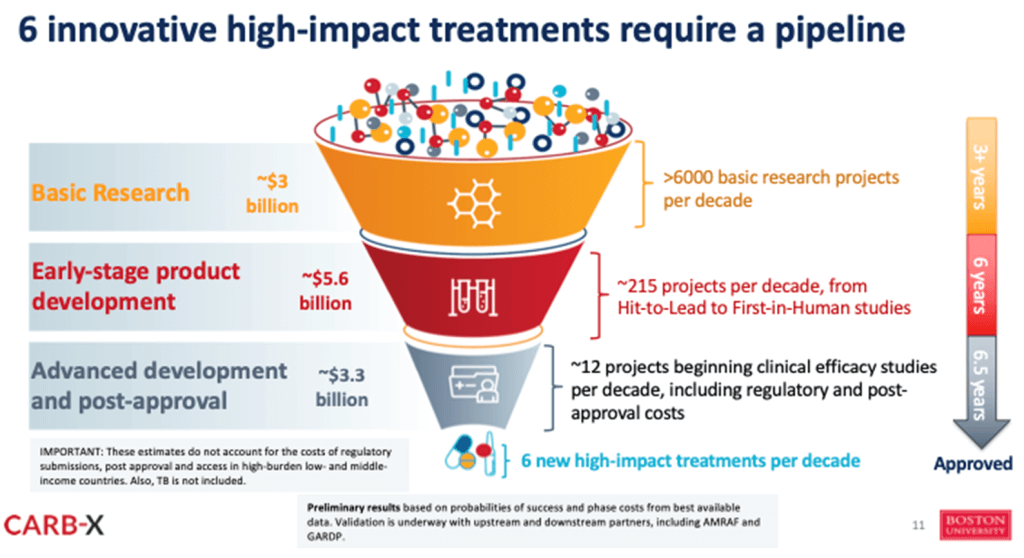
With thanks to Kevin Outterson for the slide shown just above — he used this in a talk given at the 2023 World Health Summit (Berlin, YouTube link) and tells us (personal communication) that the data are from an internal CARB-X estimate.
—
Well, enough with being depressed. As Winston Churchill said, “Success is not final, failure is not fatal: it is the courage to continue that counts.”
Our hats are off to Team GARDP for tackling this challenge as a proof of principle … Good try! It was especially interesting to see compound selection based on most current physico-chemical thinking combined with the use of a screen based on whole cell (viable, living) organisms as ways to maximize the likelihood that any resulting hits would have at least some ability to reach a target in a living bacterium.
We were also impressed to see that the authors set the bar high by focusing the search entirely on compounds with activity vs. MDR K. pneumoniae and XDR A. baumannii. Yes, these are the most difficult bacterial of all for discovery … but activity here is what would really make a difference. As said by David Burnham, “Make no little plans. They have no magic to stir men’s blood!”
Viewed from those angles and as summarized by the authors, the study met its big picture goal: “Our high-throughput phenotypic screen and selection cascade for generation of hit compounds active against multi-drug-resistant (MDR) strains of K. pneumoniae and XDR A. baumannii aims to reduce the overall cost of preclinical development by excluding NCEs at an early stage of discovery that would otherwise require significant medicinal chemistry campaigns to overcome the permeability and efflux challenges and is well documented to have had limited success.”
We contacted the authors and were told that the strategy continues to bear fruit: Laura Piddock (senior author) wrote saying, “Since completing the work described in this publication, a total of 192,000 compounds and just under 4000 bacterial natural product extracts have been tested in GARDP’s high throughput phenotypic screen. GARDP has also analysed the properties of 3.5 million compounds to help selection for HTS. Of 70 projects carried out so far, currently 22 are live in the GARDP discovery and exploratory research team.”
Exciting!
—
With gratitude towards those who keep up the hunt,
With fingers crossed for success in their efforts to expand and diversify the scope of early compound discovery, and
With all best wishes, John & Patricia
John H. Rex, MD | Chief Medical Officer, F2G Ltd. | Operating Partner, Advent Life Sciences. Follow me on Twitter: @JohnRex_NewAbx. See past newsletters and subscribe for the future: https://amr.solutions/blog/. All opinions are my own.
Patricia Bradford, PhD | Antimicrobial Development Specialists, LLC| pbradford@antimicrobialdev.com. All opinions are my own.
- BARDA’s long-running BAA (Broad Agency Announcement) for medical countermeasures (MCMs) for chemical, biological, radiological, and nuclear (CBRN) threats, pandemic influenza, and emerging infectious diseases is now BAA-23-100-SOL-00004 and offers support for both antibacterial and antifungal agents (as well as antivirals, antitoxins, diagnostics, and more). Note especially these Areas of Interest: Area 3.1 (MDR Bacteria and Biothreat Pathogens), Area 3.2 (MDR Fungal Infections), and Area 7.2 (Antibiotic Resistance Diagnostics for Priority Bacterial Pathogens). Although prior BAAs used a rolling cycle of 4 deadlines/year, the updated BAA released 26 Sep 2023 has a 5-year application period that ends 25 Sep 2028 and is open to applicants regardless of location: BARDA seeks the best science from anywhere in the world! See also this newsletter for further comments on the BAA and its areas of interest.
- HERA Invest was launched August 2023 with €100 million to support innovative EU-based SMEs in the early and late phases of clinical trials. Part of the InvestEU program supporting sustainable investment, innovation, and job creation in Europe, HERA Invest is open for application to companies developing medical countermeasures that address one of the following cross-border health threats: (i) Pathogens with pandemic or epidemic potential, (ii) Chemical, biological, radiological and nuclear (CBRN) threats originating from accidental or deliberate release, and (iii) Antimicrobial resistance (AMR). Non-dilutive venture loans covering up to 50% of investment costs are available. A closing date is not posted insofar as I can see — applications are accepted on a rolling basis; go here for more details.
- The AMR Action Fund is open on an ongoing basis to proposals for funding of Phase 2 / Phase 3 antibacterial therapeutics. Per its charter, the fund prioritizes investment in treatments that address a pathogen prioritized by the WHO, the CDC and/or other public health entities that: (i) are novel (e.g., absence of known cross-resistance, novel targets, new chemical classes, or new mechanisms of action); and/or (ii) have significant differentiated clinical utility (e.g., differentiated innovation that provides clinical value versus standard of care to prescribers and patients, such as safety/tolerability, oral formulation, different spectrum of activity); and (iii) reduce patient mortality. It is also expected that such agents would have the potential to strongly address the likely requirements for delinked Pull incentives such as the UK (NHS England) subscription pilot and the PASTEUR Act in the US. Submit queries to contact@amractionfund.com.
- INCATE (Incubator for Antibacterial Therapies in Europe) is an early-stage funding vehicle supporting innovation vs. drug-resistant bacterial infections. The fund provides advice, community, and non-dilutive funding (€10k in Stage I and up to €250k in Stage II) to support early-stage ventures in creating the evidence and building the team needed to get next-level funding. Details and contacts on their website (https://www.incate.net/).
- These things aren’t sources of funds but would help you develop funding applications
- AiCuris’ AiCubator offers incubator support to very early stage projects. Read more about it here.
- The Global AMR R&D Hub’s dynamic dashboard (link) summarizes the global clinical development pipeline, incentives for AMR R&D, and investors/investments in AMR R&D.
- Diagnostic developers would find valuable guidance in this 6-part series on in vitro diagnostic (IVD) development. Sponsored by CARB-X, C-CAMP, and FIND, it pulls together real-life insights into a succinct set of tutorials.
- In addition to the lists provided by the Global AMR R&D Hub, you might also be interested in my most current lists of R&D incentives (link) and priority pathogens (link).
John’s Top Recurring Meetings
Virtual meetings are easy to attend, but regular attendance at annual in-person events is the key to building your network and gaining deeper insight. My personal favorites for such in-person meetings are below. Of particular value for developers are the AMR Conference and the ASM-ESCMID conference. Hope to see you there!
- 27-30 April 2024 (Barcelona, Spain): 34th ECCMID, the annual meeting of the European Society for Clinical Microbiology and Infectious Diseases. Go here for details.
- 17-20 Sep 2024 (Porto, Portugal): ASM/ESCMID Joint Conference on Drug Development to Meet the Challenge of Antimicrobial Resistance. Go here for the meeting’s general website. You can’t register (yet) for the 2024 event, but save the date!
- 16-20 Oct 2024 (Los Angeles, USA): IDWeek 2024, the annual meeting of the Infectious Diseases Society of America. Save the date! More details to come!
- 25-26 February 2025 (Basel, Switzerland): The 9th AMR Conference 2025. Go here to register!
Upcoming meetings of interest to the AMR community:
- 9 Apr 2024 (virtual, 830a-10a ET): GARDP’s next REVIVE webinar entitled “Progressing a discovery project – Criteria and challenges”. Register here.
- 9 Apr 2024 (virtual, 10a-1130a ET): CDC webinar “Impacts of Antimicrobial Resistance on Cancer Care”. Click here for details and to register.
- 10-11 Apr 2024 (virtual): Sepsis Alliance AMR Conference, a 2-day conference focused on “practical technologies to manage sepsis and counteract the expanding challenge of antimicrobial resistance.” Go here for details and to register.
- 26 Apr 2024 (Barcelona, Spain): ESCMID workshop entitled “Using Data Science and Machine Learning for Infection Science: A Hands-on Introduction.” Click here to register or here for more details.
- 27-30 April 2024 (Barcelona, Spain): 34th ECCMID, the annual meeting of the European Society for Clinical Microbiology and Infectious Diseases. See Recurring Meetings list, above.
- 26-31 May 2024 (Montreal, Canada): EDAR7, the McGill AMR Centre’s 7th edition of their Environmental Dimension of Antimicrobial Resistance conference. Go here for details; final abstract deadline is 21 Dec 2023.
- 28-29 May 2024 (in person, Uppsala, Sweden): Uppsala Antibiotic Days, a broad-ranging 2-day program hosted by the Uppsala Antibiotic Center. Go here for details and to register.
- 30-31 May 2024 (face-to-face in Rockville, Maryland as well as online, 8.30-5.30p ET on 30 May, 9-2.40p on 31 May): NIAID-sponsored workshop entitled “Towards realizing the promise of adjunctive immune therapy for invasive fungal infections”. The agenda covers host immunity to invasive fungal infections, immune modulators in the context of fungal infections; and strategies for testing immune modulators as adjunctive therapy. Go here for more details and to register.
- 9-13 June 2024 (in person, Ascona, Switzerland): “New Approaches to Combat Antibiotic-Resistant Bacteria, 2nd Edition” is a Sunday-Thursday residential workshop focused on the deep biology of AMR. Sponsored by NCCR AntiResist (a Swiss National Science Foundation consortium), the scientific program has the feel of a Gordon Conference. Space is limited, so you are encouraged to apply promptly — go here for details.
- 13-17 June 2024 (Atlanta, Georgia): ASM Microbe, the annual meeting of the American Society for Microbiology. You can’t register yet, but you can go here for general details.
- 17-20 Sep 2024 (Porto, Portugal): ASM/ESCMID Joint Conference on Drug Development to Meet the Challenge of Antimicrobial Resistance. See Recurring Meetings list, above.
- 16-20 Oct 2024 (Los Angeles, USA): IDWeek 2024, the annual meeting of the Infectious Diseases Society of America. See Recurring Meetings list, above.
- 19-27 Oct 2024 (Annecy, France, residential in-person program): ICARe (Interdisciplinary Course on Antibiotics and Resistance). Now in its 8th year, Patrice Courvalin directs the program with the support of an all-star scientific committee and faculty. The resulting soup-to-nuts training covers all aspects of antimicrobials, is very intense, and routinely gets rave reviews! Seating is limited, so mark your calendars now if you are interested. Applications open in March 2024 — go here for more details.
- 4-5 Dec 2024 (in person, Washington, DC): “Fungal Dx 2024: Fungal Diagnostics in Clinical Practice” is a 2-day in-person workshop organized by ISHAM‘s Fungal Diagnostics Working Group. The program and registration links are available at https://fungaldx.com/; the agenda is comprehensive and features an all-star global list of speakers.